Permalink
Cannot retrieve contributors at this time
Name already in use
A tag already exists with the provided branch name. Many Git commands accept both tag and branch names, so creating this branch may cause unexpected behavior. Are you sure you want to create this branch?
ssHMM/README.md
Go to fileThis commit does not belong to any branch on this repository, and may belong to a fork outside of the repository.
executable file
36 lines (19 sloc)
2.72 KB
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| ## ssHMM - Sequence-structure hidden Markov model | |
| ssHMM is an RNA motif finder. It recovers sequence-structure motifs from RNA-binding protein data, such as CLIP-Seq data. | |
| ### Background | |
| RNA-binding proteins (RBPs) play a vital role in the post-transcriptional control of RNAs. They are known to recognize RNA molecules by their nucleotide sequence as well as their three-dimensional structure. ssHMM combines a hidden Markov model (HMM) with Gibbs sampling to learn the joint sequence and structure binding preferences of RBPs from high-throughput RNA-binding experiments, such as CLIP-Seq. The model can be visualized as an intuitive graph illustrating the interplay between RNA sequence and structure. | |
| ### Scope | |
| ssHMM was developed for the analysis of data from RNA-binding assays. Its aim is to help biologists to derive a binding motif for one or a number of RNA-binding proteins. ssHMM was written in Python and is a pure command-line tool. | |
| ### Output | |
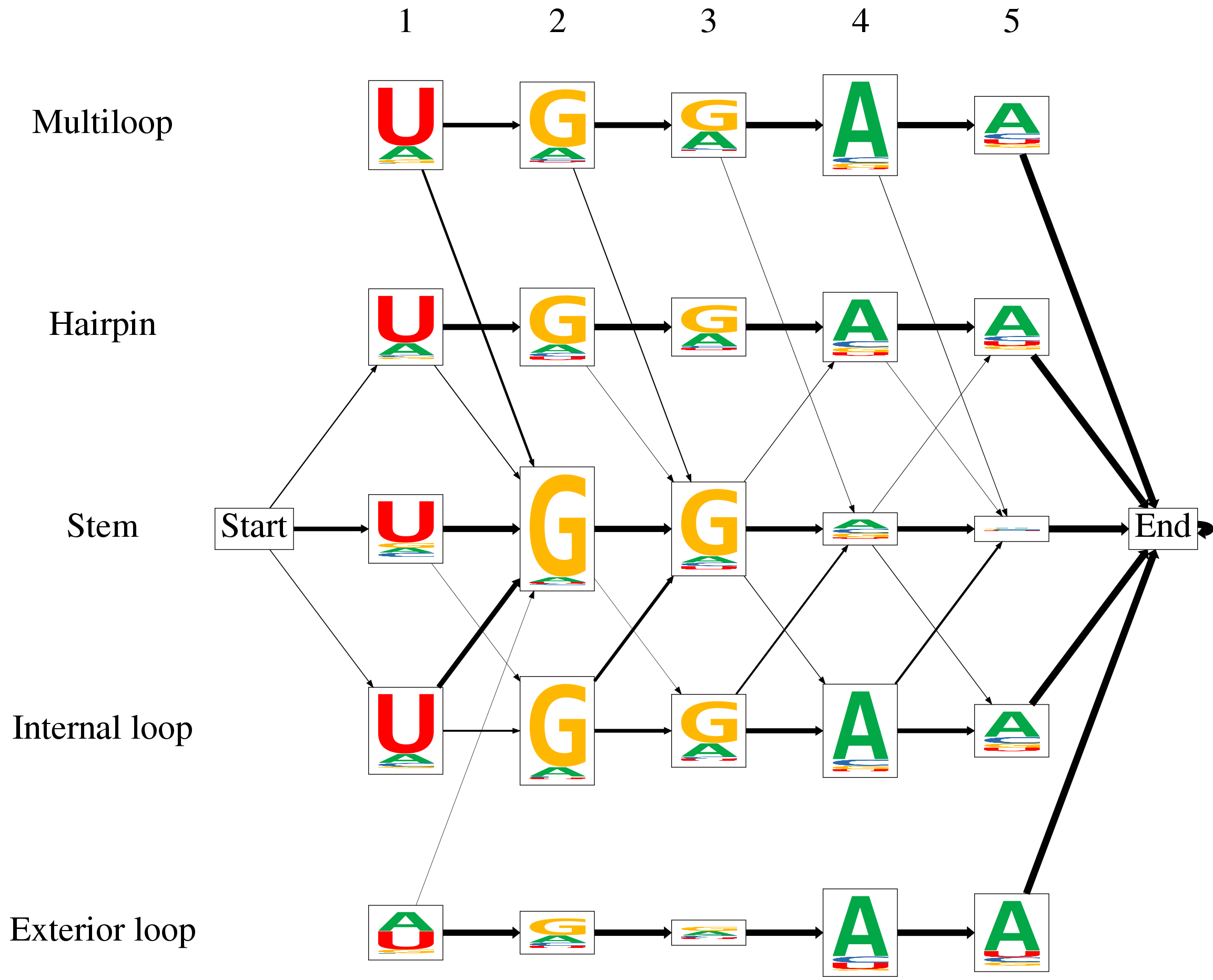
| Below, you find an example output of ssHMM. After training on an RBP dataset, the model can be visualized as a graph. Each of the HMM states, including the start and the end state, is represented by a box. The states are arranged into rows and columns where the columns represent the motif positions and the rows represent the five structural contexts of RNA. Each state's emission probabilities are visualized as a sequence logo. The relative heights of the nucleotides in a stack represent their emission probabilities. The total height of all four nucleotides is scaled according to the information content of the state's emission distribution. The transition probabilities between the states are visualized as arrows. The thicker an arrow between two states, the more likely is a transition between the two. To reduce clutter, no arrow is shown if the transition probability is less than 0.05. The RBP visualized below prefers the sequence motif UGGAA and primarily binds stem regions of RNA. | |
|  | |
| ### Documentation | |
| ssHMM can be installed manually as a Python package or (more simply) as a Docker image. Check out our documentation: http://sshmm.readthedocs.io | |
| ### Contact | |
| If you experience problems or have suggestions please contact me (see below). Alternatively, you are welcome to create issues or pull requests. | |
| Email: heller_d@molgen.mpg.de | |
| Twitter: https://twitter.com/compbiod | |
| ### Citation | |
| David Heller, Ralf Krestel, Uwe Ohler, Martin Vingron, Annalisa Marsico; ssHMM: extracting intuitive sequence-structure motifs from high-throughput RNA-binding protein data, Nucleic Acids Research, gkx756, https://doi.org/10.1093/nar/gkx756 | |
| ### License | |
| The project is licensed under the GNU General Public License. |